Author: Dániel Unyi
Link prediction is to predict whether two components in a network are likely to interact with each other. It’s a fundamental task in network science, with a wide variety of real-world applications. Examples include predicting friendship links on social media, identifying hidden communities, or discovering drug-drug interactions in pharmacology. However, current state-of-the-art algorithms are unable to scale efficiently for large graphs. My goal was to work out a scalable, accurate link prediction method by exploiting the modeling power of deep neural networks.
Introduction
Graph-based deep learning generalizes traditional methods by allowing connectivity between data points [1]. Accordingly, each data point is associated with a feature vector and an adjacency vector. Feature vectors are organized into an N×D feature matrix as usual, and adjacency vectors are organized into an N×N adjacency matrix that represents a graph (N is the number of data points and D is the number of features).
Method
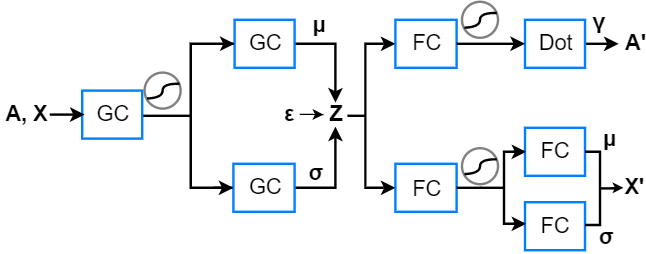
The proposed method is a variational autoencoder (VAE) [2][3] with a few modifications. Two input matrices are fed into the VAE, a feature matrix X and an adjacency matrix A. I use graph convolutional (GC) layers in the encoder to obtain the latent variables Z. I use fully connected (FC) layers in the two decoders to generate two output matrices, X’ and A’. The adjacency matrix is modelled as a set of independent Bernoulli variables: an entry represents either a link or a non-link. Predicted links are the ones which are present in A’ but not present in A. The latent variables and the feature matrix are both modelled as independent Gaussian variables.
GC layers operate in two steps. The first step is equivalent to a fully connected layer. The second step is the actual graph filtering, which can be implemented in multiple ways. The exact implementation has restrictive complexity since it requires the full diagonalization of the graph Laplacian. I applied an approximation based on the Lanczos algorithm [4]. The complexity of one Lanczos iteration is proportional to the number of links in the graph, and in my experiments, 3 iterations provided satisfying results.
Above a certain number of nodes, training in batches becomes inevitable. The basic idea could be cutting the graph into subgraphs, using an efficient clustering algorithm to keep as many links as possible. However, this approach carries two major issues. First, clustering algorithms tend to group similar nodes together, such that the distribution of a batch might be different from the distribution of the whole dataset. Second, some links are inherently removed, and the removed links are excluded from the link prediction process. To avoid these issues, I adopted the multiple stochastic clustering approach [5]. Rather than considering one cluster as a batch, I randomly choose multiple clusters. Then I form a subgraph that includes both within-cluster and between-cluster links. Hence different nodes are incorporated in a batch and there are no inherently removed links.
Results
I tested the proposed method on several real-world networks, two of which were the Reddit dataset and the DisGeNET knowledge platform. In both cases, I removed a random subset of links (15% for Reddit and 30% for DisGeNET) and trained the model on the incomplete adjacency matrix. Validation and test sets were formed from the previously removed links and the same number of randomly sampled non-links.
In the Reddit dataset, nodes represent Reddit posts committed in September 2014. If the same user commented on two posts, a link is drawn between them. Node features are GloVe CommonCrawl word vectors, containing the average embedding of the post’s title and all the post’s comments.
In the DisGeNET knowledge platform, part of the nodes represent human diseases, and the other part represents gene variants associated with them. If the occurrence of a gene variant causes disease, a link is drawn between them. Node features are node vectors embedded by node2vec.
Reddit is one of the largest graph datasets available online. Since other link prediction methods are unable to scale efficiently, the reported result can be considered as a baseline for future work. Cutting the graph into 1500 clusters and using 20 clusters per batch, a model with 2,027,520 parameters requires only 2 GB memory to train.
DisGeNET is one magnitude smaller, so in this case, I had previous baseline results. My method outperforms all of them, including the current state-of-the-art SkipGNN [6] (91.5%). I also hypothesised that some of the newly constructed links are newly discovered disease-gene associations. There’s plenty of evidence in the literature that reinforces my hypothesis. For instance, the association between liver cirrhosis and the SOD2 gene is confirmed by [7].
References
[1] Bronstein, M. M., Bruna, J., LeCun, Y., Szlam, A., & Vandergheynst, P. (2017). Geometric deep learning: going beyond Euclidean data. IEEE Signal Processing Magazine, 34(4), 18–42.
[2] Kingma, D. P., & Welling, M. (2013). Auto-encoding variational Bayes. arXiv preprint arXiv:1312.6114.
[3] Kipf, T. N., & Welling, M. (2016). Variational graph auto-encoders.
arXiv preprint arXiv:1611.07308.
[4] Tremblay, N., Gonçalves, P., & Borgnat, P. (2018). Design of graph filters and filterbanks. In Cooperative and Graph Signal Processing (pp. 299–324). Academic Press.
[5] Chiang, W. L., Liu, X., Si, S., Li, Y., Bengio, S., & Hsieh, C. J. (2019, July). Cluster-GCN: An efficient algorithm for training deep and large graph convolutional networks. In Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining (pp. 257–266).
[6] Huang, K., Xiao, C., Glass, L., Zitnik, M., & Sun, J. (2020). SkipGNN: Predicting Molecular Interactions with Skip-Graph Networks. arXiv preprint arXiv:2004.14949.
[7] Nahon, P. et al. (2009). Myeloperoxidase and superoxide dismutase 2 polymorphisms comodulate the risk of hepatocellular carcinoma and death in alcoholic cirrhosis. Hepatology, 50(5), 1484–1493.
