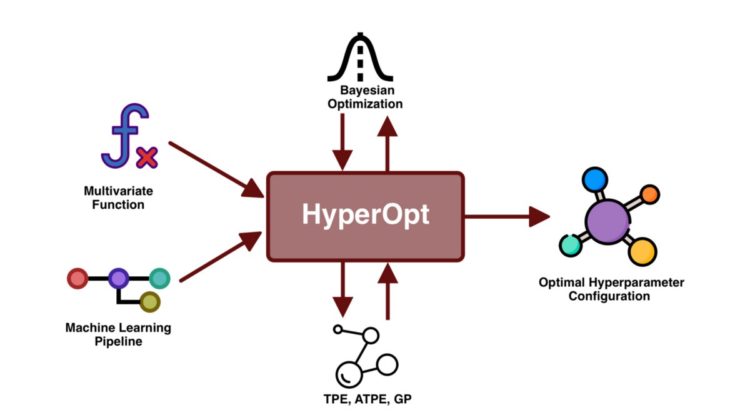
In short, HyperOpt was designed to optimize hyperparameters of one or several given functions under the paradigm of Bayesian optimization. On the other hand, HyperOpt-Sklearn was developed to optimize different components of a machine learning pipeline using HyperOpt as the core and taking various components from the scikit-learn suite. Now let’s see how we use them in practice.
Now that we understand how HyperOpt works and what its components are, let’s look at a basic implementation. For this example, we are going to use the function shown in Figure 4. As we can see, the minimum of the function is given when the value of x = -0.5. Let’s see how to find this value in HyperOpt.
HyperOpt requires 4 parameters for a basic implementation which are: the function to be optimized, the search space, the optimizer algorithm and the number of iterations. So the implementation would look like this:
As we can see, we are defining each component that HyperOpt requires to optimize a dummy function. In line 7 the definition of the function to be optimized is performed. In line 11 the definition of the search space is carried out, in this case only one search space was defined for the value of “x”, however, for functions with more than one variable, it will be required to define a search space for each variable, likewise such search space will depend on the type of function to be optimized. In this case, for didactic purposes, a search space was defined from -2 to 2. Finally, on line 18, the class that hosts the optimization process is initialized. Such function receives as parameters the function to be optimized, the search space, the optimization algorithm (in this case it is Tree-structured of Parzen Estimators) and the number of iterations. When executing the previous code snippet we obtain the value of “x” that optimizes the function:
Optimal value of x: {'x': -0.5000806428004325}
The previous implementation is a basic example of how HyperOpt works and what its main components are. The optimization of more elaborate functions will require the adequate definition of the search space as well as the optimizer. HyperOpt provides a set of search space initializers which you can find here.
Great, we have already seen how HyperOpt works in a basic implementation, now let’s see how HyperOpt-Sklearn works for Machine Learning pipeline optimization.
The way to implement HyperOpt-Sklearn is quite similar to HyperOpt. Since HyperOpt-Sklearn is focused on optimizing machine learning pipelines, the 3 essential parameters that are required are: the type of preprocessor, the machine learning model (i.e. classifier or regressor) and the optimizer. It is important to mention that each of these three basic elements are customizable according to the needs of each problem.
The preprocessors adapted in HyperOpt Sklearn are: PCA, TfidfVectorizer, StandardScalar, MinMaxScalar, Normalizer, OneHotEncoder. The classifiers adapted in HyperOpt Sklearn are: SVC, LinearSVC KNeightborsClassifier. RandomForestClassifier, ExtraTreesClassifier SGDClassifier, MultinomialNB, BernoulliRBM, ColumnKMeans.
For the purposes of this blog, let’s look at two basic implementations of HyperOpt-Sklearn in a classification problem. In this example we will work with the well known breast cancer dataset.
As we can see, in line 22 we are defining the classifier that will be implemented, in this case the instruction is to search over all the classifiers defined by HyperOpt-Sklearn (in practice this is not recommended due to the computation time needed for the optimization, since this is a practical example, doing a full search is not a determining factor). In line 23 the type of transformation that the data will receive is defined, in this case the instruction to use the complete suite of transformers implemented by HyperOpt-Sklearn (as you can guess, only those that fit the dataset are tested, e.g. text transformers are not applied to a numeric dataset). In line 24 the optimizer is defined, in this case it is TPE. The rest of the lines determine the number of iterations and the time limit for each evaluation.
When executing the code snippet 2 we obtain
Train score: 0.9723618090452262
Test score: 0.9824561403508771
The optimal configuration is:
{'learner': ExtraTreesClassifier(max_features=None, min_samples_leaf=9, n_estimators=19, n_jobs=1, random_state=3, verbose=False), 'preprocs': (MinMaxScaler(feature_range=(-1.0, 1.0)),), 'ex_preprocs': ()}
Well, we have obtained an optimal configuration by doing a search across the entire spectrum that HyperOpt-Sklearn covers for a classification problem. Now let’s see how we would narrow down the search space by using a specific classifier.
For this example we will also use the breast cancer dataset. However, this time we will use a single classifier of which we will try to optimize each of its parameters by defining a search space for each one.
In this example we are using SGD as a classifier for which we want to optimize the loss parameter as well as the alpha value. As we can see, in line 23 we are defining a search space for loss, such a search space is defined by three different values (hinge, log, huber) with a probability value that is considered when selecting one of these three values. On the other hand, in line 29 we are defining the search space for the alpha value, in this case a log function is implemented which is bounded by a lower and upper limit. Finally, on line 33, the class that will host the optimization process is defined. The parameters it receives are the classifier (with their respective parameters and search spaces), the optimizer, the number of iterations, and the designated time for each evaluation.
When executing the code snippet 3we obtain
Train score: 0.9522613065326633
Test score: 0.9473684210526315
The optimal configuration found is:
{'learner': SGDClassifier(alpha=0.08612797536101766, class_weight='balanced', eta0=6.478871110431366e-05, l1_ratio=0.20803307323675568, learning_rate='invscaling', loss='log', max_iter=18547873.0, n_jobs=1, power_t=0.1770890191026292, random_state=0, tol=0.000332542442869532, verbose=False),
'preprocs': (PCA(n_components=8),), 'ex_preprocs': ()}
HyperOpt-Sklearn configurations and customizations will always depend on the type of problem to be solved, the data types as well as the available computing power.
In this blog we saw what HyperOpt is, its purpose, how it works and what its main components are. Likewise, we saw one of the main extensions of HyperOpt which is HyperOpt-Sklearn, its components and how it works.
HyperOpt is an alternative for the optimization of hyperparameters, either in specific functions or optimizing pipelines of machine learning. One of the great advantages of HyperOpt is the implementation of Bayesian optimization with specific adaptations, which makes HyperOpt a tool to consider for tuning hyperparameters.
[1] Tuning the hyper-parameters of an estimator
[2] TPOT: Pipelines Optimization with Genetic Algorithms
[3] A Tutorial on Bayesian Optimization
[5] Algorithms for Hyper-Parameter Optimization
[6] Hyperopt-Sklearn: Automatic Hyperparameter Configuration for Scikit-Learn
