This article explains stratified cross-validation and it’s implementation in Python using Scikit-Learn. This article assumes the reader to have a working knowledge of cross-validation in machine learning.
What is stratified sampling?
Before diving deep into stratified cross-validation, it is important to know about stratified sampling. Stratified sampling is a sampling technique where the samples are selected in the same proportion (by dividing the population into groups called ‘strata’ based on a characteristic) as they appear in the population. For example, if the population of interest has 30% male and 70% female subjects, then we divide the population into two (‘male’ and ‘female’) groups and choose 30% of the sample from the ‘male’ group and ‘70%’ of the sample from the ‘female’ group.
How is stratified sampling related to cross-validation?
Implementing the concept of stratified sampling in cross-validation ensures the training and test sets have the same proportion of the feature of interest as in the original dataset. Doing this with the target variable ensures that the cross-validation result is a close approximation of generalization error.
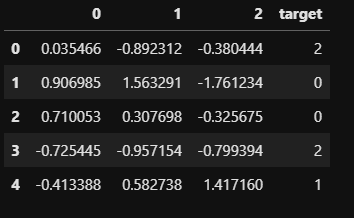
Before proceeding further, we’ll generate a synthetic classification dataset with 500 records, three features and three classes. The dataset is generated using ‘make_classification’ function of Scikit-Learn.
import pandas as pd
import numpy as np
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split, StratifiedKFold, StratifiedShuffleSplit, KFoldmake_class = make_classification(n_samples=500,n_features=3,n_redundant=0,n_informative=2,n_classes=3,n_clusters_per_class=1,random_state=11)data = pd.DataFrame(make_class[0],columns=range(make_class[0].shape[1]))data['target'] = make_class[1]data.head()
Implementing hold-out cross-validation without stratified sampling
Hold-out cross validation is implemented using the ‘train_test_split’ function of Scikit-Learn. The implementation is shown below. The function returns training set and test set.
train_df,test_df = train_test_split(data,test_size=0.2,random_state=11)print(f'PROPORTION OF TARGET IN THE ORIGINAL DATAn{data["target"].value_counts() / len(data)}nn'+
f'PROPORTION OF TARGET IN THE TRAINING SETn{train_df["target"].value_counts() / len(train_df)}nn'+
f'PROPORTION OF TARGET IN THE TEST SETn{test_df["target"].value_counts() / len(test_df)}')
Since, we haven’t used stratified sampling, we can see that the proportion of the target variable varies hugely among the original dataset, training set and test set.
Implementing hold-out cross-validation with stratified sampling
We’ll implement hold-out cross-validation with stratified sampling such that the training and the test sets have same proportion of the target variable. This can be achieved by setting the ‘stratify’ argument of ‘train_test_split’ to the characteristic of interest (target variable, in this case). It need not necessarily be the target variable, it can even be an input variable which you want to have the same proportion in the training and test sets.
train_df,test_df = train_test_split(data,test_size=0.2,stratify=data['target'],random_state=11)print(f'PROPORTION OF TARGET IN THE ORIGINAL DATAn{data["target"].value_counts() / len(data)}nn'+
f'PROPORTION OF TARGET IN THE TRAINING SETn{train_df["target"].value_counts() / len(train_df)}nn'+
f'PROPORTION OF TARGET IN THE TEST SETn{test_df["target"].value_counts() / len(test_df)}')
Using stratified sampling, the proportion of the target variable is pretty much the same across the original data, training set and test set.
Implementing k-fold cross-validation without stratified sampling
K-fold cross-validation splits the data into ‘k’ portions. In each of ‘k’ iterations, one portion is used as the test set, while the remaining portions are used for training. Using the ‘KFold’ function of Scikit-Learn, we’ll implement 3-fold cross-validation without stratified sampling.
kfold = KFold(n_splits=3,random_state=11,shuffle=True)splits = kfold.split(data,data['target']) # each split has a train indexes and test indexes pairprint(f'PROPORTION OF TARGET IN THE ORIGINAL DATAn{data["target"].value_counts() / len(data)}nn')
for n,(train_index,test_index) in enumerate(splits):
print(f'SPLIT NO {n+1}nTRAINING SET SIZE: {np.round(len(train_index) / (len(train_index)+len(test_index)),2)}'+
f' TEST SET SIZE: {np.round(len(test_index) / (len(train_index)+len(test_index)),2)}nPROPORTION OF TARGET IN THE TRAINING SETn'+
f'{data.iloc[test_index,3].value_counts() / len(data.iloc[test_index,3])}nPROPORTION OF TARGET IN THE TEST SETn'+
f'{data.iloc[train_index,3].value_counts() / len(data.iloc[train_index,3])}nn')
We can see that the proportion of the target variable is inconsistent among the original data, training data and test data across splits.
Implementing k-fold cross-validation with stratified sampling
Stratified sampling can be implemented with k-fold cross-validation using the ‘StratifiedKFold’ function of Scikit-Learn. The implementation is shown below.
kfold = StratifiedKFold(n_splits=3,shuffle=True,random_state=11)#data['target'] IS THE VARIABLE USED FOR STRATIFIED SAMPLING.
splits = kfold.split(data,data['target'])print(f'PROPORTION OF TARGET IN THE ORIGINAL DATAn{data["target"].value_counts() / len(data)}nn')
for n,(train_index,test_index) in enumerate(splits):
print(f'SPLIT NO {n+1}nTRAINING SET SIZE: {np.round(len(train_index) / (len(train_index)+len(test_index)),2)}'+
f' TEST SET SIZE: {np.round(len(test_index) / (len(train_index)+len(test_index)),2)}nPROPORTION OF TARGET IN THE TRAINING SETn'+
f'{data.iloc[test_index,3].value_counts() / len(data.iloc[test_index,3])}nPROPORTION OF TARGET IN THE TEST SETn'+
f'{data.iloc[train_index,3].value_counts() / len(data.iloc[train_index,3])}nn')
In the above results we can see that the proportion of the target variable is pretty much consistent across the original data, training set and test set in all the three splits.
Cross-validation implemented using stratified sampling ensures that the proportion of feature of interest is same across the original data, training set and the test set. This ensures that no value is over/under represented in the training and test sets, which gives a more accurate estimate of performance/error.
